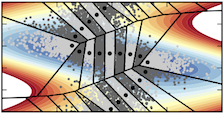
Dickson Periodic Potential¶
Driven Brownian particle on a 2D periodic surface based on the potential and parameterization found in:
A. Dickson, A. Warmflash, and A. R. Dinner, “Nonequilibrium umbrella sampling in spaces of many order parameters,” J Chem Phys 130, 074104 (2009). http://dx.doi.org/10.1063/1.3070677
and
A. Dickson, A. Warmflash, and A. R. Dinner, “Erratum: ”Nonequilibrium umbrella sampling in spaces of many order pa- rameters” [J. Chem. Phys. 130, 074104 (2009)],” J Chem Phys 136, 229901 (2012). http://dx.doi.org/10.1063/1.4729744
Running conventional simulations¶
To run the brute force simulations, use the script, bin/run_bruteforce.py
$ python bin/run_bruteforce.py -h
usage: run_bruteforce.py [-h] [-s NSIMS] -c CONFIG_FILE [CONFIG_FILE ...] -n
[NAME [NAME ...]] [-w NWORKERS]
[--sid_offset SID_OFFSET]
Bruteforce run script
optional arguments:
-h, --help show this help message and exit
-s NSIMS number of simulations to run
-c CONFIG_FILE [CONFIG_FILE ...] yaml config file name
-n [NAME [NAME ...]] simulation name to run
-w NWORKERS number of cores to use
--sid_offset SID_OFFSET offset for numbering simulations
This and other scripts uses python’s multiprocessing module to run multiple independent simulations on multiple cores simultaneously. The number of workers defaults to the number of cores available, although this can be specified via the -w flag. The --sid_offset command line argument offsets the simulation identifier.
The commands used for the paper were:
$ python bin/run_bruteforce.py -s 20 -c configs/bruteforce_run_config.yaml -n bruteforce_common -w 6
and:
$ python bin/run_bruteforce.py -s 20 -c configs/bruteforce_run_config.yaml -n bruteforce_rare -w 6
to run the brute force simulations for the parameterization of the potential where transitions are common and rare, respectively. Since these and other simulations can take a long time to complete, it is advisable to use nohup.
Running Weighted Ensemble simulations¶
To run the Weighted Ensemble simulations, use the script, bin/run_we.py
$ python bin/run_we.py -h
usage: run_we.py [-h] [-s NSIMS] -c CONFIG_FILE [CONFIG_FILE ...] -n
[NAME [NAME ...]] [-p [PROTOCOLS [PROTOCOLS ...]]]
[-w NWORKERS] [--sid_offset SID_OFFSET] [--no-run]
WEST run script
optional arguments:
-h, --help show this help message and exit
-s NSIMS number of simulations to run
-c CONFIG_FILE [CONFIG_FILE ...]
yaml config file name
-n [NAME [NAME ...]] simulation name to run
-p [PROTOCOLS [PROTOCOLS ...]]
protocols to run; by default run all
-w NWORKERS number of cores to use
--sid_offset SID_OFFSET
offset for numbering simulations
--no-run Only setup simulations but do not run them
This tool is similar to run_bruteforce.py, but adds two additional command line arguments; The -p flag specifies which protocols to run (DEFAULT: run all), and --no-run sets up all of the simulation directories and scripts, but does not launch the actual simulations. The protocols are specified in the yaml configuration file in configs and the are selected by the name variable.
The commands used for the paper were:
$ python bin/run_we.py -s 10 -c configs/we_common_run_config.yaml -n we_common -w 6
and:
$ python bin/run_we.py -s 10 -c configs/we_rare_run_config.yaml -n we_rare -w 6
to run the Weighted Ensemble simulations for the parameterization of the potential where transitions are common and rare, respectively.
Analyzing data¶
To analyze the Weighted Ensemble simulations, use the script, bin/run_we_analysis.py
$ python bin/run_we_analysis.py -h
usage: run_we_analysis.py [-h] -c CONFIG_FILE [CONFIG_FILE ...]
[-n [NAME [NAME ...]]] [-w NWORKERS] -s
{calculate_distribution,get_pcoords,get_strings,get_nreplicas,all}
optional arguments:
-h, --help show this help message and exit
-c CONFIG_FILE [CONFIG_FILE ...]
yaml config file name
-n [NAME [NAME ...]] simulation name to run; by default run all
-w NWORKERS number of cores to use
-s {calculate_distribution,get_pcoords,get_strings,get_nreplicas,all}
analysis script to run
This script runs the analysis tools found in the analysis directory. The commands used for the paper were:
$ python bin/run_we_analysis.py -c configs/we_common_run_config.yaml -n we_common -w 3 -s all
and:
$ python bin/run_we_analysis.py -c configs/we_rare_run_config.yaml -n we_rare -w 3 -s all
These scripts generate a number of .h5 files in we_common/analysis/{sim_id} and we_rare/analysis/{sim_id} containing the calculated quantities used to create the figures.
Generating figures¶
To generate the figures that appear in the manuscript after running all of the simulations and the scripts that analyze them, run:
$ cd generate_figures
$ python distributions.py
$ python error_plot.py
$ python potential_string.py
This will create one .eps file per figure.